HTG EdgeSeq Chemistry
Together with next-generation sequencing (NGS) instrumentation, our HTG EdgeSeq chemistry allows for multiplexed, quantitative molecular profiling using a 5 µm FFPE tissue section or 15 µl of plasma/serum.
Features
- Extraction-free
- Measure the expression levels of a few hundred genes to whole mRNA and miRNA transcriptome in a single reaction.
- Requires very little sample input
HTG EdgeSeq Chemistry Overview
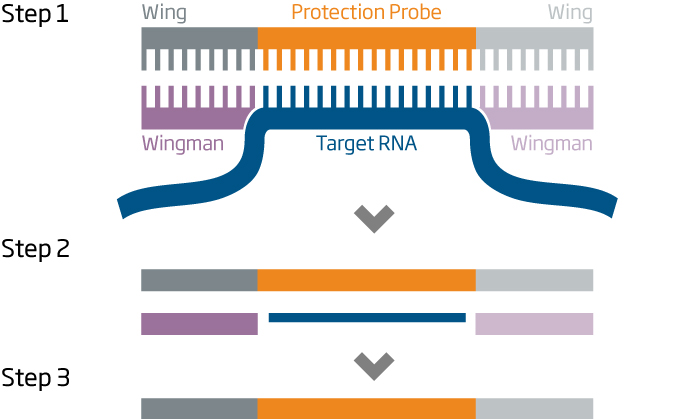
HTG EdgeSeq assays employ nuclease protection to measure RNA. Target-specific protection probes with universal priming sites (wings) hybridize to target transcripts and wingman (1). S1 nuclease is added to digest non-hybridized probes and RNA, resulting in a 1-to-1 ratios of probes and target RNAs (2).
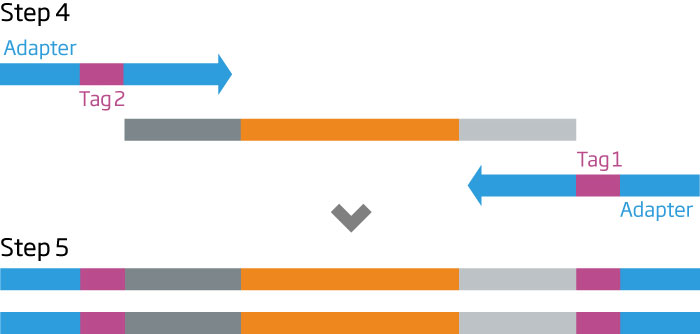
After S1 nucleuse inactivation (3), the remaining probes are amplified with primers carrying sequencing adaptors and molecular barcodes (4). The resulting PCR products are purified, quantitated, and combined to make a sequencing ready library (5). HTG EdgeSeq assays do not require processing steps such as reverse transcription, adenylation, or adaptor ligation.
Steps 1 - 3 are fully automated on the HTG EdgeSeq system.
Sample Types
Our chemistry supports a wide variety of sample types, including formalin-fixed, paraffin-embedded (FFPE) samples.
Gene Expression Panels
Our expertly curated gene expression panels leverage the power and sensitivity of next-generation sequencing, allowing you to measure the expression levels of a few hundred genes to whole mRNA and miRNA transcriptome. Our breakthrough technology generates results from clinically relevant samples, such as formalin-fixed, paraffin-embedded (FFPE) tissue, to support biomarker discovery, drug targeting, and diagnostics development.
HTG Research Use Only (RUO ) Panels
HTG EdgeSeq Research Use Only (RUO) Panels
Learn More
Learn more about the HTG EdgeSeq Chemistry.
Page last updated May 23, 2022
Ready to get started?
HTG is focused on delivering exceptional molecular profiling products and services. Contact us to find out how our unique technology can benefit you.