HTG EdgeSeq Oncology Biomarker Panel
The HTG EdgeSeq Oncology Biomarker Panel (OBP) is a systems biology pathway analysis tool that profiles samples to identify therapeutic targets and drug response markers. The OBP assay utilizes our automated HTG EdgeSeq system coupled with the sensitivity and dynamic range of next-generation sequencing (NGS)-based detection, and can be performed in your lab using the HTG EdgeSeq system, from our VERI/O lab as a service, or at one of the Qualified Service Provider (QSP) sites.
For Research Use Only. Not for Use in Diagnostic Procedures.
Features and Benefits
- Simultaneous, quantitative detection of 2,549 genes associated with tumor biology
- Extraction-free: reduce extraction-associated data bias and sample loss
- Reliable, reproducible performance from a single formalin-fixed, paraffin-embedded (FFPE) needle core biopsy slide
- Gain insights faster: results in as little as 36 hours
- Simplified data analysis: raw numerical data output provided in Excel format
Sample Requirements
| Sample Type | Recommended Sample Input |
|---|---|
Larger sample input amounts must be diluted. Please consult the assay package insert and HTG EdgeSeq System User Manual for more details. |
|
| Extracted RNA from FFPE | 35 ng |
| PAXgene | 500 µl |
| FFPE Tissue | One 5 µm section |
| Cells | ≥3,000 cells |
| Extracted RNA from Frozen | 10 - 35 ng |
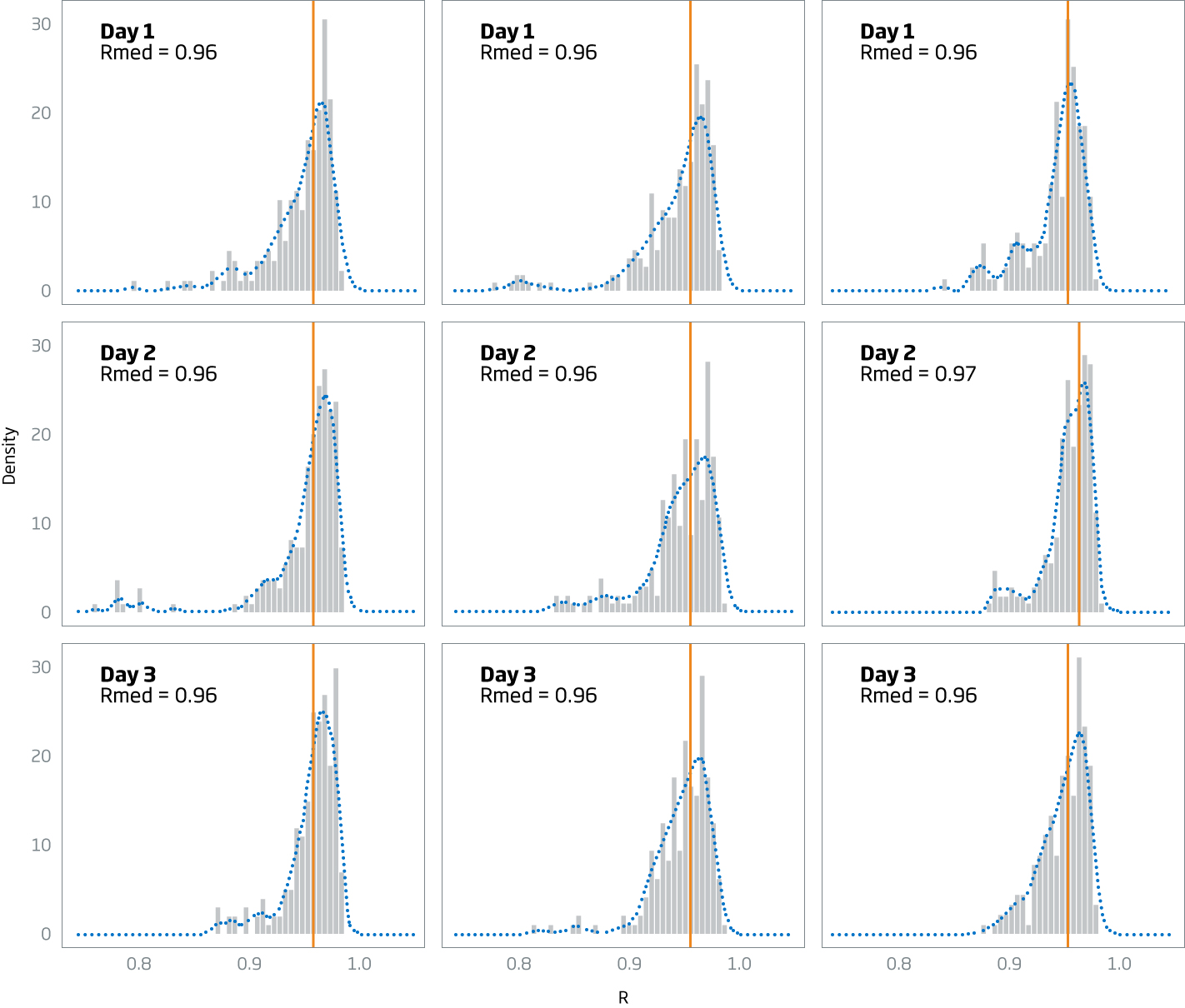
Performance
Pearson correlations for all replicate pairs within each condition are summarized in the table below. The vertical line in the histogram represents the median correlation for all 225 Pearson correlation pairs.
| Processor | Day 1 | Day 2 | Day 3 | Overall |
|---|---|---|---|---|
| P23 | 0.96 | 0.96 | 0.96 | 0.96 |
| P31 | 0.96 | 0.96 | 0.96 | 0.96 |
| P33 | 0.96 | 0.97 | 0.96 | 0.96 |
| Overall | 0.96 | 0.96 | 0.96 | 0.96 |
Applications
- Assess FFPE tumor samples for potential biomarkers
- Subtype tumors based on cellular origin, treatment response, or aggressiveness
- Develop predictive and prognostic gene signatures
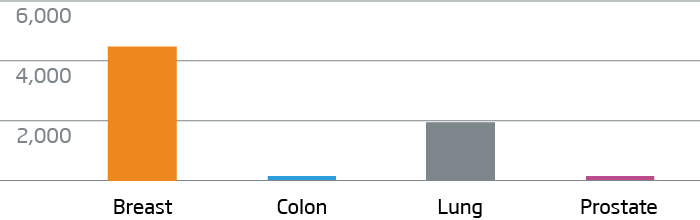
Breast, colon, lung, and prostate FFPE tumor tissue were processed with the HTG EdgeSeq Oncology Biomarker Panel. Data are displayed as normalized counts per million (CPM).
KLK3 (PSA) - Prostate Cancer Marker
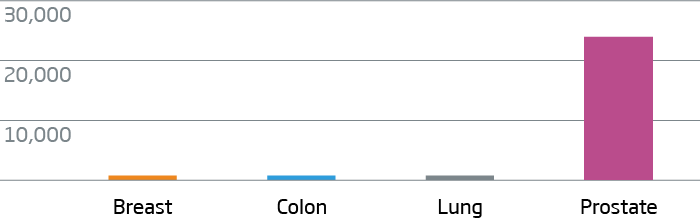
CDX2 - Colorectal Cancer Marker
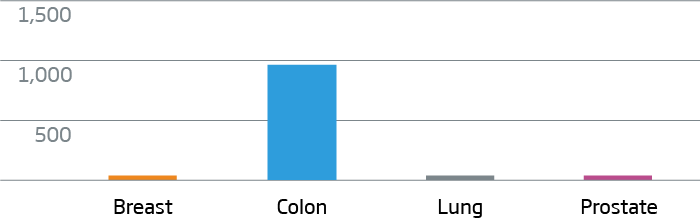
KRT7 - Breast and Lung Cancer Marker
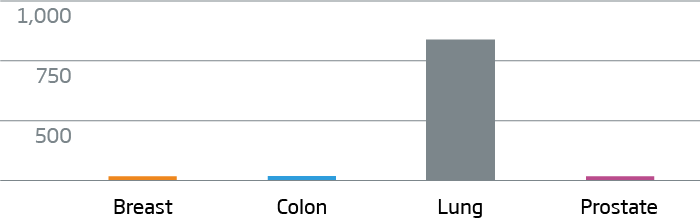
NKX2-1 (TTF-1) - Lung Cancer Marker
Ordering Information
When placing an order, please specify the catalog number. The HTG EdgeSeq Oncology Biomarker Panel is compatible with both Illumina sequencers and Thermo Fisher Ion Torrent sequencers.
Kit Configurations for use with Illumina Sequencers
916-002-208 HTG EdgeSeq Oncology Biomarker Panel (2x8)
916-002-008 HTG EdgeSeq Oncology Biomarker Panel (4x8)
916-002-224 HTG EdgeSeq Oncology Biomarker Panel (1x24)
916-002-024 HTG EdgeSeq Oncology Biomarker Panel (4x24)
916-002-096 HTG EdgeSeq Oncology Biomarker Panel (1x96)
Kit Configurations for use with Thermo Fisher Scientific Ion Torrent Sequencers
916-002-308 HTG EdgeSeq Oncology Biomarker Panel (2x8)
916-002-108 HTG EdgeSeq Oncology Biomarker Panel (4x8)
916-002-324 HTG EdgeSeq Oncology Biomarker Panel (1x24)
916-002-124 HTG EdgeSeq Oncology Biomarker Panel (4x24)
916-002-196 HTG EdgeSeq Oncology Biomarker Panel (1X96)
For Research Use Only. Not for Use in Diagnostic Procedures.
Resources and Publications
Learn more about the HTG EdgeSeq Oncology Biomarker Panel
Integrated Molecular Profiling of Genitourinary Tumors: Applications for Personalized Radiotherapy
View Webinar
FFPE Tissue and the HTG EdgeSeq Oncology Biomarker Panel
View Webinar
Expression-Based Profiling of Challenging FFPE Tumor Samples
View Webinar
HTG EdgeSeq Oncology Biomarker Panel Product Sheet
Download pdf 967KB
Select and download available HTG EdgeSeq Oncology Biomarker Panel genes
View Link
Publications
Novel EHE PDX model used for drug sensitivity
View External Link
IOA-244 is a Non–ATP-competitive, Highly Selective, Tolerable PI3K Delta Inhibitor That Targets Solid Tumors and Breaks Immune Tolerance
View External Link
Abstract P3-05-09: LAG3+ Tumor Infiltrating Lymphocytes Predict Outcome in Treatment Naïve Triple Negative Breast Carcinoma
View External Link
Abstract PD17-05: Development and Validation of a Composite Biomarker Predictive of Palbociclib + Endocrine Treatment Benefit in Early Breast Cancer: PENELOPE-B and PALLAS Trials
View External Link
Abstract P5-02-17: Prognostic and predictive role of RBsig and CCNE1/RB1 gene-expression signatures in patients with advanced breast cancer treated with palbociclib in combination with endocrine therapy in the PALOMA-2 and 3 trials
View External Link
53P Characterization of the non-ATP competitive PI3Kdelta inhibitor IOA-244 in lymphoma models: From single agent to combination screen and clinical investigation
View External Link
Pharmacodynamic activity of BMS-986156, a glucocorticoid-induced TNF receptor-related protein agonist, alone or in combination with nivolumab in patients with advanced solid tumors
View External Link
CCNE1 and PLK1 mediates resistance to palbociclib in HR+/HER2- metastatic breast cancer
View External Link
Reliability of gene-expression profiling from tumor biopsy for refining neoadjuvant strategies in patients with head and neck squamous cell carcinoma
View External Link
Use of distinct molecular signatures of appendiceal cancer subtypes to assess biomarker development
View External Link
Platinum-Based Regimens Are Active in Advanced Pediatric-Type Rhabdomyosarcoma in Adults and Depending on HMGB1 Expression
View External Link
SABCS 2022: HER2-06: Outcome analysis of HER2-zero or HER2-low hormone receptor-positive (HR+) breast cancer patients - characterization of the molecular phenotype in combination with molecular subtyping
Download pdf 1.0MB
View External Link
Clinical implications of the intrinsic molecular subtypes in hormone receptor-positive and HER2-negative metastatic breast cancer
View External Link
SABCS 2022: PD4-02: Spatial and temporal heterogeneity of predictive and prognostic signatures in triple-negative breast cancer treated with neoadjuvant combination immune-chemotherapy
Download pdf 942KB
View External Link
SABCS 2022: PD17-06: Immunohistochemical Markers and Determinants of Clinical Response in the Penelope-B Trial
Download pdf 939KB
View External Link
SABCS 2022: PD-17-05: Development and Validation of a Composite Biomarker Predictive of Palbociclib + Endocrine Treatment Benefit in Early Breast Cancer: PENELOPE-B and PALLAS Trials.
Download pdf 964KB
View External Link
Distinct immune signature predicts progression of vestibular schwannoma and unveils a possible viral etiology
View External Link
Immunologically active phenotype by gene expression profiling is associated with clinical benefit from PD-1/PD-L1 inhibitors in real-world head and neck and lung cancer patients
View External Link
Pan-cancer gene expression analysis of tissue microarray using EdgeSeq oncology biomarker panel and a cross-comparison with HER2 and HER3 immunohistochemical analysis
View External Link
Mutational And Gene Expression Profiling Of Blastic Plasmacytoid Dendritic Cell Neoplasm Reveals Two Distinct Age-Dependent Subgroups And A Role For Clonal Hematopoiesis In Elderly Patients
View External Link
NSABP FC-6: Surgical conversion rate in colorectal cancer patients with unresectable, KRAS wild-type liver metastases receiving mFOLFOX7 plus cetuximab
View External Link
NKG2A and HLA-E define an alternative immune checkpoint axis in bladder cancer
View External Link
Datasets for gene expression profiles of head and neck squamous cell carcinoma and lung cancer treated or not by PD1/PD-L1 inhibitors
View External Link
Comparative biomarker analysis of PALOMA-2/3 trials for palbociclib
View External Link
Prognostic Value of EZH2 Expression for Immunotherapy-based Schemes in Advanced Soft-tissue Sarcoma
Download pdf 2.1MB
Biomarkers for Response to Immunotherapy in Triple-Negative Breast Cancer - Differences Between Survival and pCR Biomarkers
Download pdf 674KB
P-481 Characterization of the gene expression profile of breast cancer tumours in patients undergoing ovarian stimulation for fertility preservation purposes
View External Link
OTHR-43. Composition of Cell-Free miRNA in Cerebrospinal Fluid and Plasma as a Monitoring Tool for Pediatric Brain Tumors
View External Link
Genetic and phenotypic attributes of splenic marginal zone lymphoma
View External Link
Abstract PD2-07: Impact of using cross-platform gene expression profiling technologies and computational methods for intrinsic breast cancer subtyping in PALOMA-2 and PALLET
View External Link
Oropharyngeal cancer outcomes correlate with p16 status, multinucleation and immune infiltration
View External Link
Abstract PD2-04: Molecular plasticity of luminal breast cancer and response to CDK 4/6 inhibition - The biomarker program of the PENELOPE-B trial investigating post-neoadjuvant palbociclib
View External Link
Biological processes associated with the efficacy of sunitinib plus nivolumab in soft tissue sarcoma: correlative studies of the IMMUNOSARC trial of Spanish (GEIS) and Italian (ISG) sarcoma groups - poster
Download pdf 1.0MB
Tomentosin a Sesquiterpene Lactone Induces Antiproliferative and Proapoptotic Effects in Human Burkitt Lymphoma by Deregulation of Anti- and Pro-Apoptotic Genes
View External Link
The pulmonary pathology of COVID-19. Virchows Arch (2021). https://doi.org/10.1007/s00428-021-03053-1 2.21.2021 FFPE
Download pdf 6.6MB
HTG EdgeSeq technology offers a competitive alternative to RNA-Seq with equivalent performance and distinct advantages.
Download pdf 613KB
Enapotamab vedotin, an AXL-specific antibody-drug conjugate, shows preclinical antitumor activity in non-small cell lung cancer.
Download pdf 3.8MB
Reliable Gene Expression Profiling from Small and Hematoxylin and Eosin-Stained Clinical Formalin-Fixed, Paraffin-Embedded Specimens Using the HTG EdgeSeq Platform
View External Link
Gene expression and genetic variants in Parkinson's disease (PD) genes to predict outcome in metastatic colorectal cancer (mCRC): Data from FIRE-3 phase III trial.
View External Link
Baseline tumor-immune signatures associated with response to bempegaldesleukin (NKTR-214) and nivolumab
View External Link
Gene expression comparison between primary triple-negative breast cancer and matched axillary lymph node metastasis
Download pdf 1.2MB
View External Link
Association of an Inflammatory Gene Signature with CD8 Expression by Immunohistochemistry (IHC) in Multiple Tumor Types.
Download pdf 606KB
View External Link
Predicting Response to Neoadjuvant Therapy in Oesophageal Adenocarcinoma Pre-Treatment Biopsies
Download pdf 1.5MB
EMT- and stroma-related gene expression and resistance to PD-1 blockade in urothelial cancer
Download pdf 2.0MB
Randomized phase II neoadjuvant study (GepartNuevo) to investigate the addition of durvalumab to a taxane-anthracycline containing chemotherapy in triple negative breast cancer (TNBC)
Download pdf 2.8MB
View External Link
Correlation between gene expression and prognostic biomarkers in small cell bladder cancer (SCBC).
Download ppt 1.2MB
View External Link
Effect of rilimogene galvacirepvec/rilimogene glafolivec on intra/peritumoral immune infiltrate in patients with localized prostate cancer undergoing radical prostatectomy.
View External Link
A biologic basis for locoregional failure in patients with oral cavity cancers
View External Link
Integrative omics to detect bacteremia in patients with febrile neutropenia
Download pdf 3.1MB
Prognostic value of CD56, ASCL1, and other biomarkers in small cell bladder cancer (SCBC).
View External Link
PQR309 is a novel dual P13K/mTOR inhibitor with pre-clinical anti-tumor activity in lymphomas as a single agent and in combination therapy.
View External Link
Phase II study of dovitinib in patients (Pts) progressing on anti-vascular endothelial growth factor (VEGF) therapy
Download pdf 848KB
Association of gene signature to identify molecular subtypes with clinical outcomes of 1st-line cetuximab (cet) treatment for metastatic colorectal cancer (mCRC)
View External Link
Evaluation and selection of a non-PCR based technology for improved gene expression profiling from clinical formalin-fixed, paraffin-embedded samples
Download pdf 1.0MB
View External Link
Gene expression profiling of transplant formalin-fixed, paraffin-embedded biopsies: comparison of a custom TaqMan low density array and quantitative nuclease-protection assay
View External Link
Learn More
For further information about the HTG EdgeSeq Oncology Biomarker Panel, please fill out the registration form below. Or call us at (877) 507-3259.
Page last updated June 29, 2022